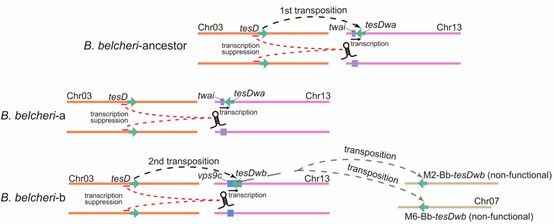
Abstract: Sex determination systems display striking evolutionary flexibility, yet the mechanisms underlying their transitions remain poorly understood. Using newly generated genome assemblies, we investigated the evolving sex-determining system in the amphioxus Branchiostoma belcheri. We identified two female-specific sex-determining regions (SDRs) on chromosome 13, both derived from independent transpositions of the autosomal gene tesD, which shows testis-specific expression in amphioxus species. CRISPR/Cas9 knockout experiments in Branchiostoma floridae confirmed that tesD functions as a male-determination gene, with loss of function producing an all-female phenotype. In B. belcheri, the older SDR (tesDwa) inserted into the coding region of twai, while the younger SDR (tesDwb), flanked by active Zator-1 transposons, inserted into the 3' UTR of vps9c and later translocated to autosomes in ~10% of individuals. Transcriptomic analyses revealed that W-linked tesDwa and tesDwb produce antisense long non-coding RNAs that likely suppress tesD transcription in females, whereas autosomal tesDwb is not expressed and appears non-functional. The insertion sites and co-transcription with host genes suggest promoter hijacking. Together, these findings demonstrate that recurrent transpositions can generate new functional SDRs that coexist with older ones, driving dynamic turnover of sex determination in B. belcheri.
Link: https://www.nature.com/articles/s41467-026-68322-6