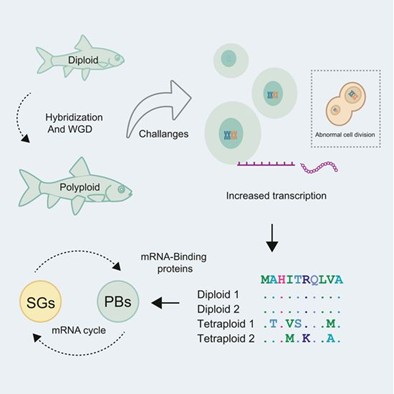
Abstract: Polyploidy, the duplication of the entire genome, is a major force in vertebrate evolution, yet how new polyploid species survive and prosper remains unclear. Here, we present a chromosome-level genome of the cyprinid Spinibarbus caldwelli, whose allotetraploid event is dated to ∼4 million years ago based on transposable element distributions in its two subgenomes. Genomic analyses reveal unique patterns of homoeologous exchanges between the two subgenomes, indicating a gradual evolutionary process prior to their suppression. Furthermore, we identify a striking pattern of accelerated evolution of RNA-binding proteins across independently evolved tetraploid cyprinids. Functional assays of one such protein, Tia1, demonstrate that the tetraploid orthologs may be more efficient at stress granule disassembly than those of diploid relatives. These findings suggest that improved cellular stress management, particularly in RNA processing, might be a key adaptation that has enabled the evolutionary success of polyploid cyprinids.
Link: https://www.cell.com/cell-reports/fulltext/S2211-1247(25)01407-X